Hybrid speciation in Cottus

Cottus from the Dutch lowlands and Belgium (C. perifretum) as well as from tributaries to the River Rhine (C. rhenanus,) have come into secondary contact and given rise to invasive Cottus, a lineage of hybrid origin. Invasive Cottus (center, top individual) have colonized and spread within the lower River Rhine (left) while C. rhenanus (center, bottom individual) are restricted to smaller tributaries as for example the Bröl in North Rhine Westphalia (right) (Pictures: I. Steinmann, A. Hartl).
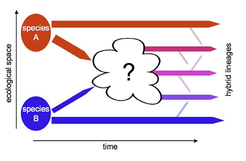
Homoploid hybrid speciation can be explained, if hybrid separate spatially and ecologically from their parents. However, intrinsic genetic conflicts and selective forces that affect in the emerging hybrid lineages are poorly understood.

We have set up aquaria in a climate controlled chamber. Here the temperature and daylight cycle can be adjusted and follow the conditions in nature. This represents a prerequisite to successfully breed Cottus in captivity.


